Genes and Mendel's Third Law
From the last post you’ll know that meiosis results in four n cells from one 2n cell. These n cells are known as gametes or sex cells — males make sperm sex cells and females make egg sex cells.
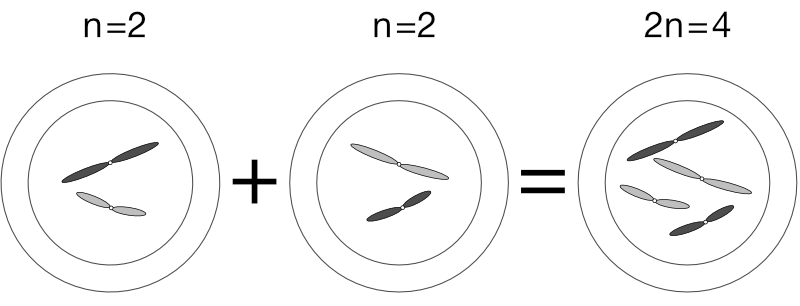
Still another term for an n cell is haploid cell. When a haploid sperm fertilises a haploid egg, the two nuclei merge to form a 2n, or diploid, cell. The haploid cells in the diagram below have two distinct chromosomes each (n = 2). All four chromosomes end up in the fertilised egg, but because the four are really two homologous pairs, we say 2n = 4:
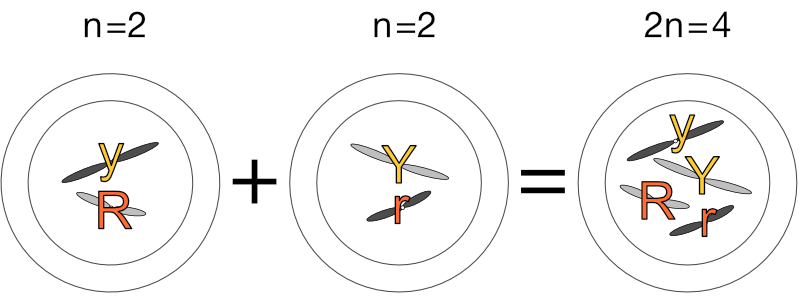
Let’s add our pea genes from the previous post to make a very simplified diploid pea cell:
This RrYy diploid cell now carries the genes for round (’R') pea shape, wrinkled pea shape (’r'), yellow seed colour (’Y') and green seed colour (’y'). Should it undergo mitosis and develop into an adult pea plant, it will only ever produce round and yellow seeds despite carrying the other genes, for round is always dominant to wrinkled, and yellow is always dominant to green.
Though this plant will only ever have round yellow seeds, it is heterozygous for both seed shape and seed colour. Thus via Mendel’s first law of segregation and his second law of independent assortment covered here at the cellular level, that plant will produce sex cells of RY, Ry, rY and ry combinations.
Mendel knew from his pea experiments that ‘R’ and ‘Y’ are dominant over ‘r’ and ‘y’ respectively, and derived his third law of dominance from these observations. He knew that a dominant character would express itself whether in the presence of a recessive one or not, but did not know the underlying reasons for this.
To explain what dominance actually is, let’s backtrack a bit.
An organism’s entire genome (its complete set of DNA, or complete set of genetic material) is spread over its full complement of chromosomes. A dog’s genome is spread over 78 chromosomes for example. You’ll know from Meiosis that chromosomes in a diploid cell exist as homologous pairs. A dog would thus have 39 homologous pairs, with one homologue of each pair having come from the mother and the other having come from the father.
Geneticists designate an order and number to the homologous pairs of species’ genomes. The largest chromosome pair is usually ‘1′, the second largest ‘2′ and so on. The very last pair is always the sex chromosomes and not numbered, but rather called X and Y in mammals or Z and W in birds.
Each homologous pair carries specific genes. In dogs, a gene for brown coat colour is found only on their chromosome 11, while a gene linked to blue eye colour is on their chromosome 18. The gene for pea seed colour is on a pea’s chromosome 1, while the gene for seed shape is on chromosome 7.
A pea has two ‘1′ chromosomes, and thus has two copies of the seed colour gene, one on each homolog. If these copies are identical (eg both are ‘Y’ or ‘y’), we say the plant is homozygous for that trait. If the copies are different (eg one is ‘Y’ and one is ‘y’), the plant is heterozygous for that trait.
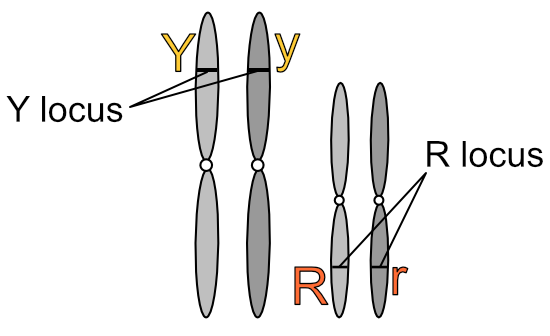
Not only are specific genes found on specific chromosomes, they are also found at specific locations on those chromosomes. A gene location is called a locus (plural: loci). Bearing in mind that the following is illustrative only, we could represent our pea genes like so:
The locus is named for the dominant form of the gene that resides there. One homologue in the above diagram carries the ‘Y’ version of the gene at the Y locus, while the other carries the ‘y’ version of the same gene. Similarly for ‘R’ and ‘r’ at the R locus. People refer to ‘dominant’ and ‘recessive’ genes — and I’ve done the same in this blog to simplify things where appropriate — but it is more correct to refer to dominant and recessive versions of a particular gene. And like everything in biology (!) these different versions of the same gene have their own name: alleles. In the diagram above, ‘Y’ and ‘y’ are the alleles for seed colour at the Y locus, and ‘R’ and ‘r’ are the alleles for seed shape at the R locus.
Though a pea plant may carry both the ‘Y’ and ‘y’ allele, something about the ‘Y’ allele dominates, as if the ‘y’ were never there. What makes a particular allele dominant?
It may surprise you to learn that a gene is nothing more than a set of instructions for making a particular protein. It may surprise you even more to learn that most enzymes and some hormones are proteins or peptides (protein subunits)! Thus proteins are crucial to the functioning of an organism.
It follows that a dominant allele is simply a piece of genetic code that makes a protein which ‘out-competes’ the protein coded by the recessive allele. In some cases the recessive allele codes for a ‘broken’ (non-functioning) protein and the dominant protein ‘wins’ by default. This is actually the case with the Y allele in pea plants.
A plant with yellow peas produces a protein which switches on a particular set of genes in the peas. That set of genes in turn produces proteins which destroy chlorophyll. Chlorophyll is a green pigment, thus destroying it leaves the peas a yellow colour. A plant with green peas is homozygous recessive — it has two copies of the ‘r’ allele. The ‘r’ allele codes for a broken protein: it is defective and cannot switch on the chlorophyll-destroying genes. With no ‘R’ allele to produce a functioning protein, the chlorophyll is undisturbed in the pea, and the pea stays green.
This example of one allele dominating over another is called complete dominance, and is classic Mendelian inheritance. The heterozygous phenotype is indistinguishable from the homozygous phenotype. Examples abound in the animal world too, with simply-inherited traits typically expressed this way. The polled (hornless) allele in cattle is completely dominant to the horned allele. The suri fleece type in alpacas is completely dominant to the huacaya fleece type. Black feathers in chickens are dominant to red feathers.
Mendel was very lucky to have chosen seven pea traits that not only happened to mostly reside on separate chromosomes, but which also followed complete dominance. More so when you know that a pea plant has 2n = 14, ie he chose seven genes on five out of a possible seven chromosomes!
As the field of genetics advanced, scientists discovered that not all inheritance was as simple or predictable as Mendel’s observations. These forms of inheritance became known as non-Mendelian inheritance, and to know and understand these emphasises not only how random inheritance truly is, but how lucky Mendel was!
Up next: non-Mendelian inheritance!
Leave a comment