We know from Mendel’s experiments that each parent contains two copies of a ‘factor’ (gene). Let’s call these two copies a gene pair.
We know from Mendel’s Laws that these gene pairs segregate and independently sort themselves inside each parent in some way, such that each parent transfers just one copy of a gene pair to its offspring. The offspring in turn ends up with its own two copies of that gene, one from each parent, and the cycle repeats.
But what is that ‘way’? What ensures an offspring ends up with a pair of genes itself, and not a lone copy, or pairs of pairs, or any other number other than two copies?
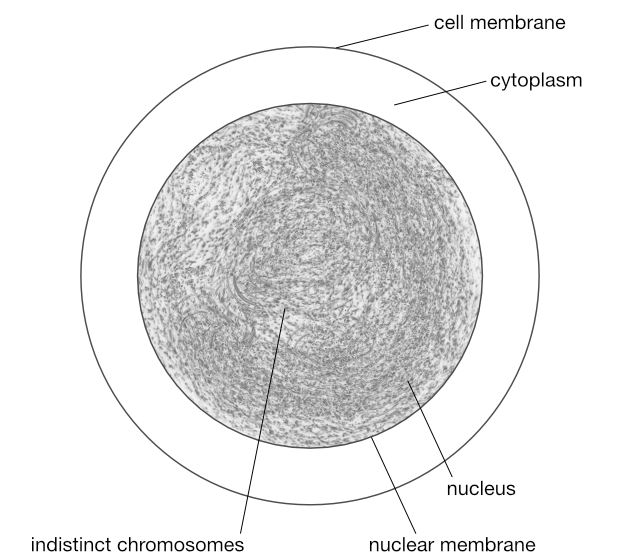
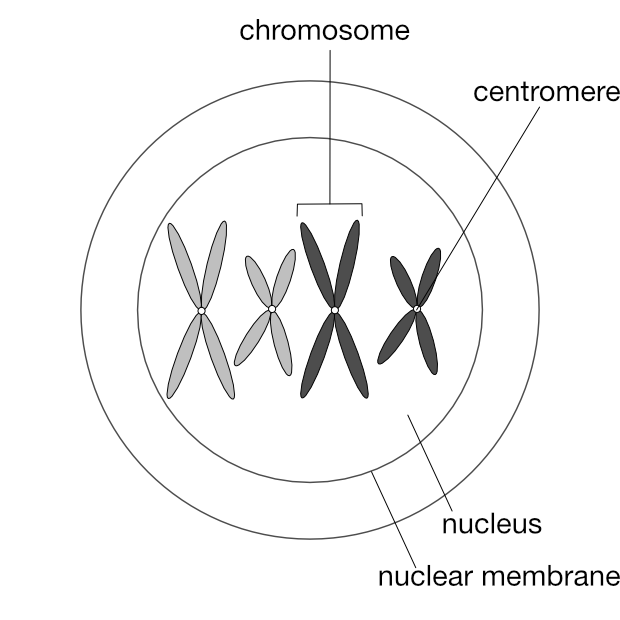
You’d know that an animal is made up of cells. Each cell contains a nucleus (there are exceptions such as the red blood cells of most mammals), and inside this nucleus is the entire genetic material of that animal spread amongst multiple strands of deoxyribonucleic acid (DNA). A DNA strand is a very long molecule containing many genes along its length. A strand of DNA is packaged into a structure called a chromosome.
The number of chromosomes differs from species to species. Humans have 46 chromosomes, while dogs have 78, as do chickens. Cattle and goats have 60. There’s no real significance to this, it’s just fun to know!
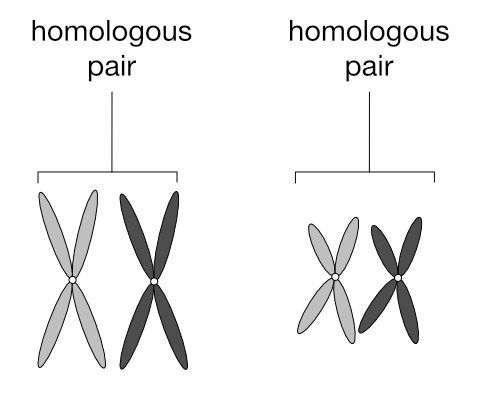
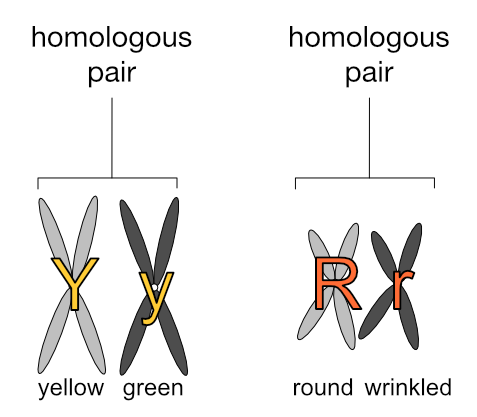
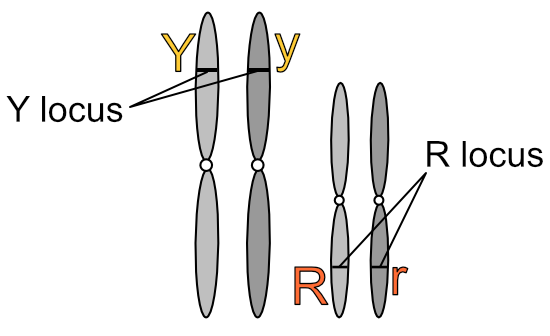
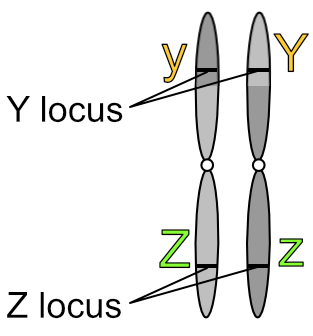
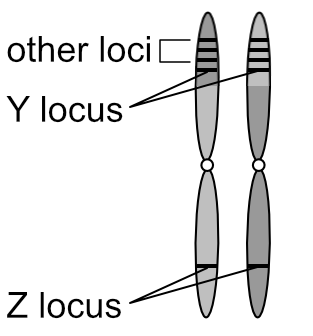
What’s more meaningful to know is that chromosomes exist in pairs. This is why genes also exist in pairs — one of each gene pair resides on one of each chromosome pair.
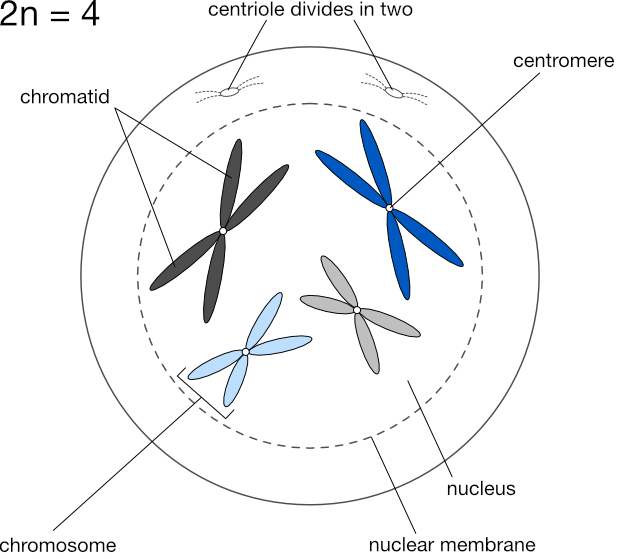
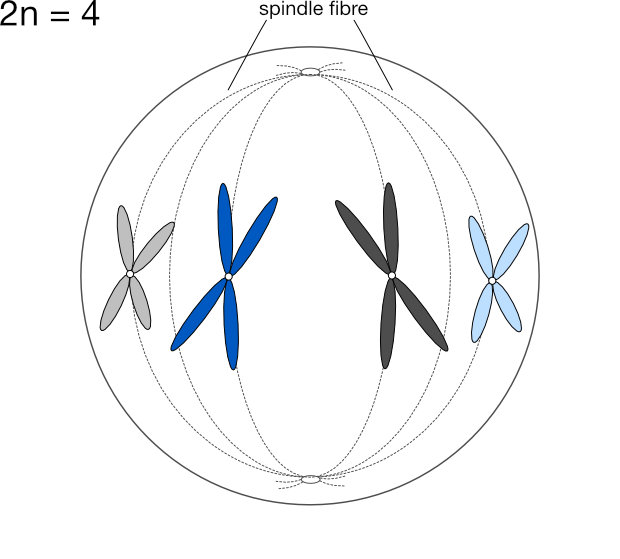
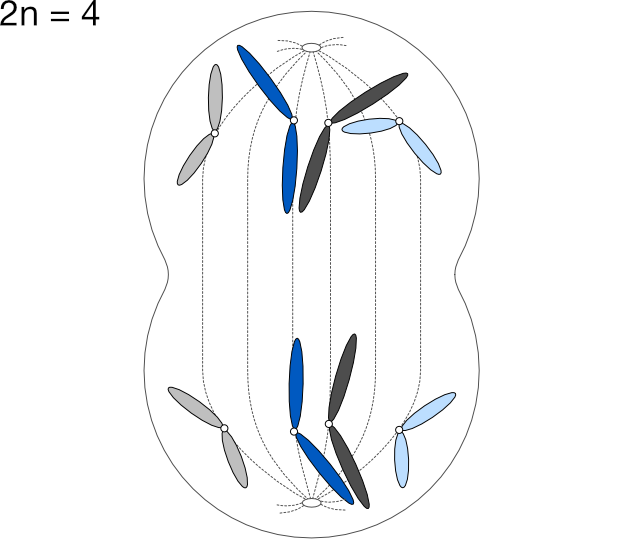
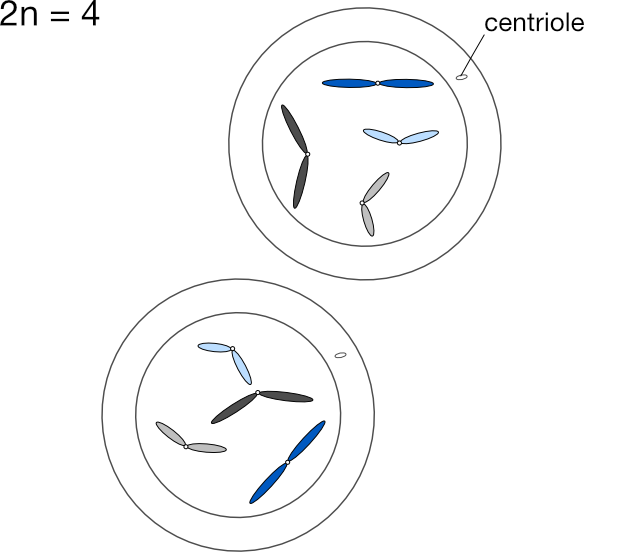
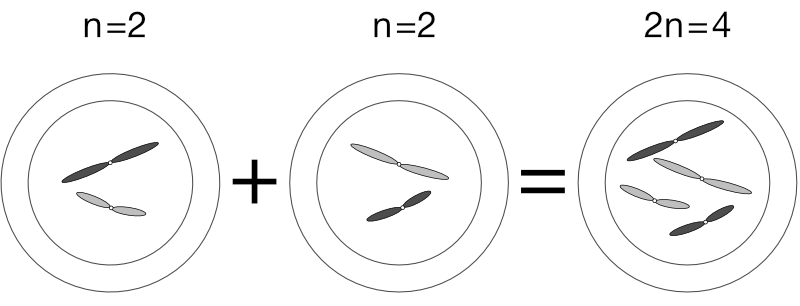
We express the number of chromosomes in an animal as 2n, where n is the number of pairs, which varies from species to species. Dogs have 78 chromosomes total, so 2n=78, meaning n, the number of pairs, is 39. Humans have n=23 pairs of chromosomes.
Every cell with a nucleus has a 2n complement of chromosomes. These cells group themselves into specialised populations to form tissues and organs. Muscle cells make up a muscle, liver cells make up a liver, kidney cells make up a kidney, and so on. Cells in these tissues and organs continually replicate themselves to replace others that die off: new muscle cells replace old muscle cells, liver cells replace liver cells, etc.
Cells replicate themselves by dividing into two identical copies with a full set of chromosomes each: for example, a 2n kidney cell produces two 2n daughter kidney cells, each of which will in turn produce two 2n daughter kidney cells of their own.This process is called mitosis, and I’ll elaborate more on this in the next post.
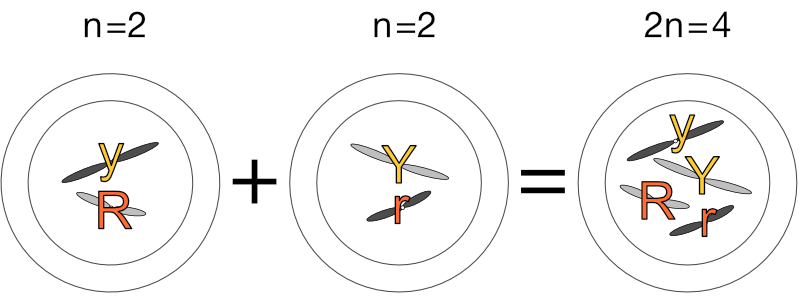
I said above that each cell in an animal has a 2n complement of chromosomes. This is true of all cells that make up the tissues and organs, with one sole exception: the sex cells produced by the sex organs. The eggs produced by the ovaries in females and the sperm produced by the testes in males have an n complement of chromosomes. In other words, a sex cell (also known as a gamete) contains just one copy of each chromosome pair, and thus one copy of each gene pair.
When a sperm fertilises an egg, the n complement of the sperm joins with the n complement of the egg, conferring a full 2n complement to the developing embryo. This is how a parent donates just one gene copy and why offspring inherit two.
This process — the production of gametes — is called meiosis, and I’ll go into far more detail on this after covering mitosis.
But how does this apply to Mendel’s first two Laws?
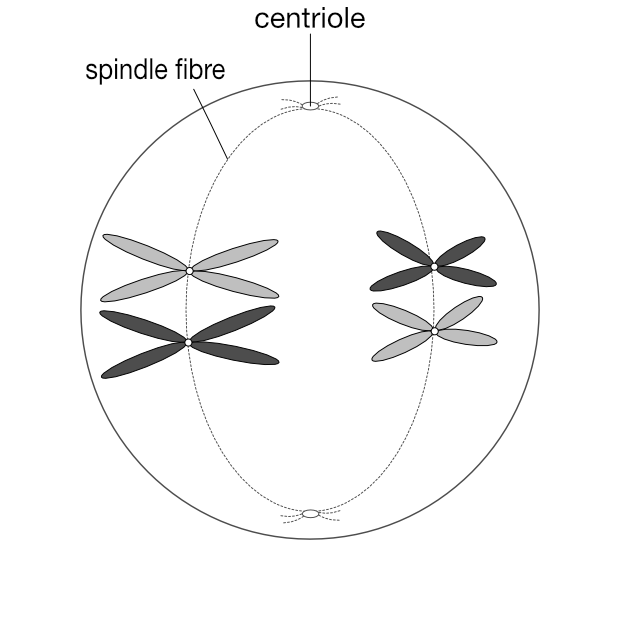
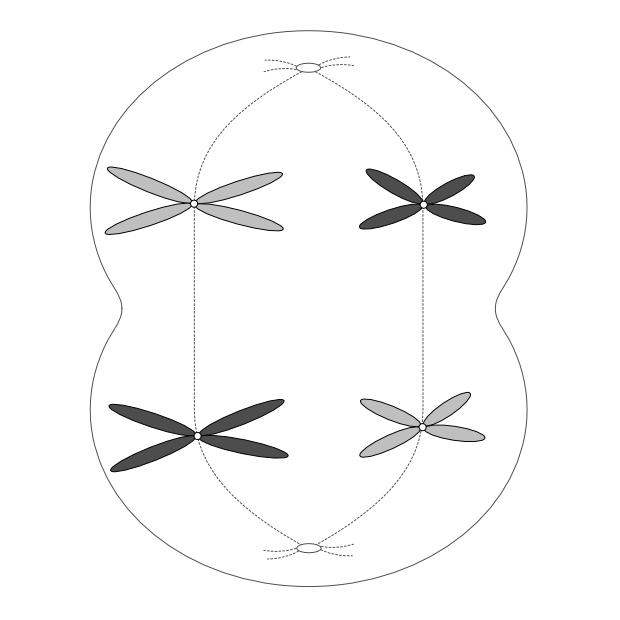
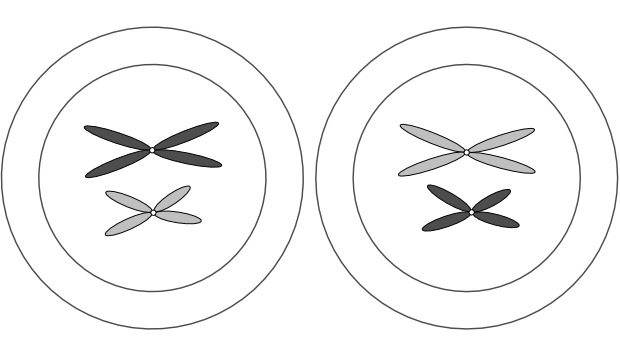
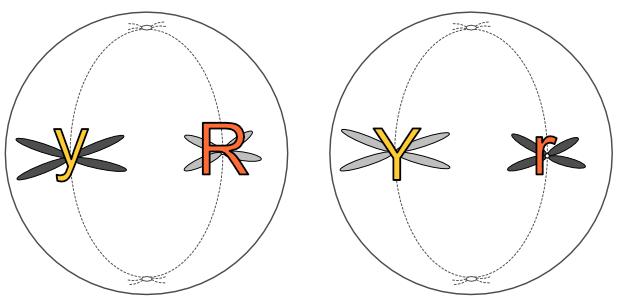
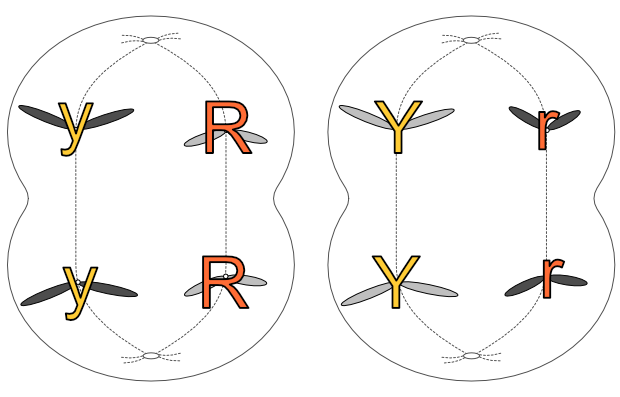
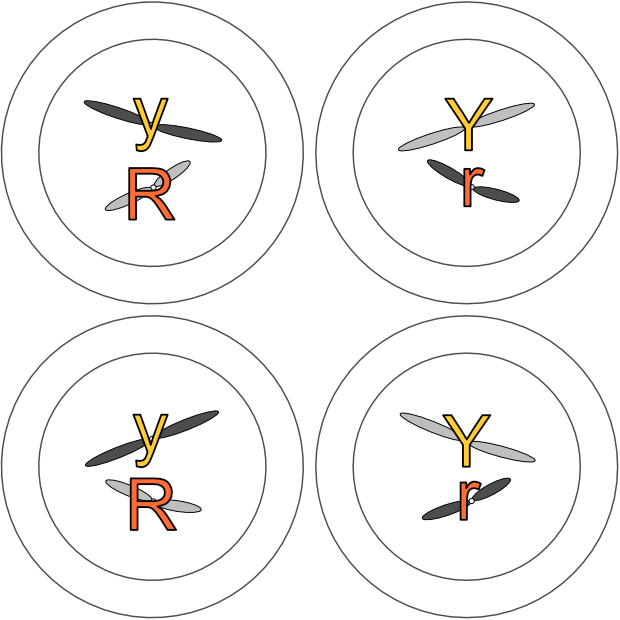
Gametes (eggs and sperm) have an n complement, but they actually arise from 2n precursors. But instead of producing two 2n cells, these precursors divide into four n cells. Unlike mitosis, where two 2n cells arise from one, in meiosis four n gametes arise from one 2n cell. Each gamete has just one copy of every original chromosome pair.
This is Mendel’s First Law, the law of segregation (of genes). One 2n cell produces four n gametes, each of which contains just one of each chromosome pair, which in turn contains one copy of each gene associated with that chromosome. The chromosome pairs — and thus the genes on those chromosomes — separate out into four distinct gametes when a 2n cell splits into four n cells.
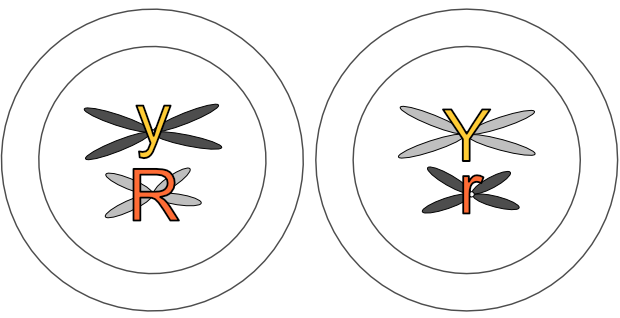
Any one of those chromosome pairs in the original 2n cell could end up in a gamete with any other one of the other chromosome pairs. Imagine a very simple 2n cell with just three chromosome pairs, named 1a and 1b, 2a and 2b, and 3a and 3b. After meiosis, we could end up with a gamete containing chromosome 1a, chromosome 2b and chromosome 3b. Or 1b, 2b and 3a. Or some other combination. (There would be eight possible combinations just from these three pairs alone — I’ll explain the maths in a later post!)
This is Mendel’s Second Law, the law of independent assortment (of genes). There is nothing determining which chromosome of any pair (and its genes) ends up with any other chromosome from any other pair in a gamete. And thus nothing determining which copy of a gene pair ends up with any other copy of a different gene pair.
This admittedly was a very brief coverage of a rather involved subject: please regard it as an introduction to set the scene. The next few weeks will be far more in depth, and with diagrams and a bit of maths to boot!