You’d know from the previous post that regular body cells such as kidney cells or liver cells undergo mitosis to make identical 2n copies of themselves that regenerate the organs and body as older cells die.
Meiosis has a different purpose. It is a specialised and more complicated form of cell division restricted to the sex organs. Through meiosis, ovaries make egg cells and testes make sperm cells containing half the number of chromosomes (half the genetic content of the parent). Meiosis makes n cells from 2n cells, and this occurs only in the sex organs.
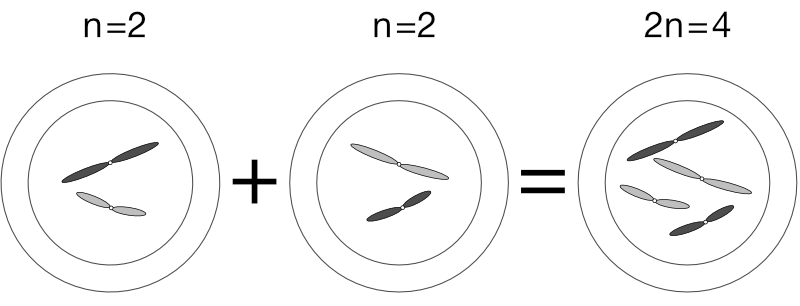
When an n sperm fertilises an n egg, its genetic material enters the egg’s nucleus and the egg acquires a full number (n + n = 2n) of chromosomes. With this complete set of genetic material, half from each parent, the fertilised egg then divides via regular mitosis, and again, and again, until after many, many divisions and a lot of complicated processes, a new and complete organism forms. In time that organism will produce its own eggs or sperm via meiosis to continue the cycle.
We’ll now step through the entire process of meiosis — and knowing how cells divide through mitosis will make this much easier to follow, as you will see!
Meiosis comprises two stages of division, Phase 1 and Phase 2. Each phase is broken down into interphase, prophase, metaphase, anaphase and telophase as with mitosis.
Phase 1
The first division. At the end of this stage the cell has divided, but the chromosomes have not separated. The following images will help explain.
Interphase 1
The chromosomes replicate themselves as they would in mitosis.
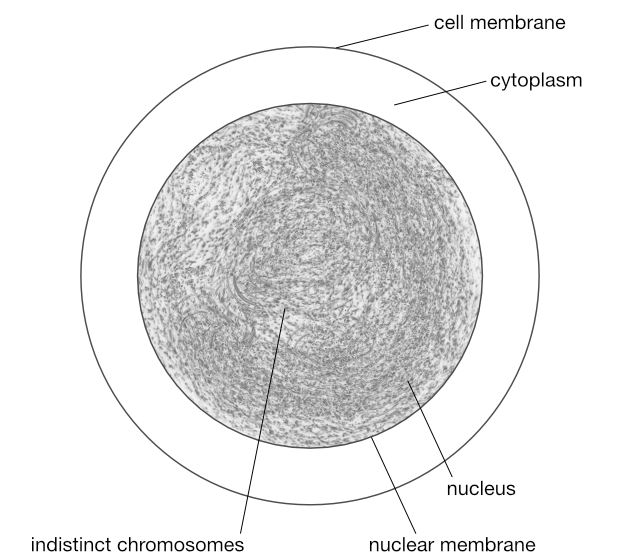
© Optimate Group Pty Ltd
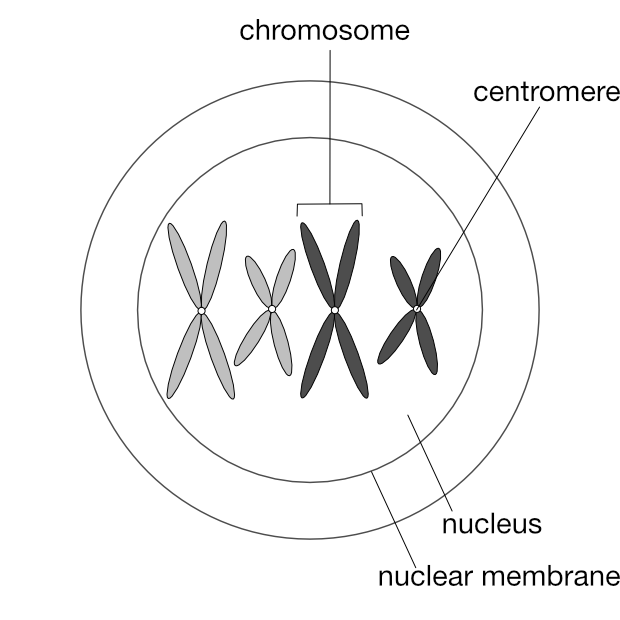
Prophase 1
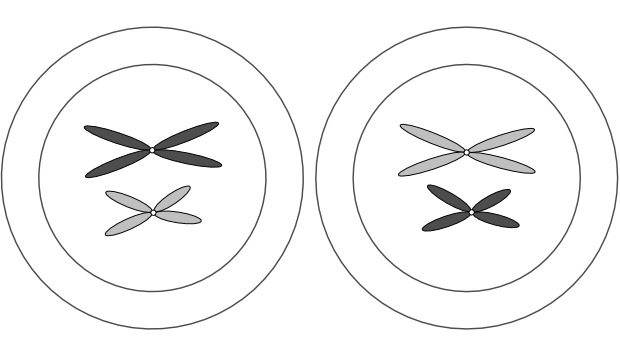
Again as in mitosis, the chromosomes are visible if stained, and appear as duplicates joined at a centromere.
Remember how chromosomes exist as pairs?
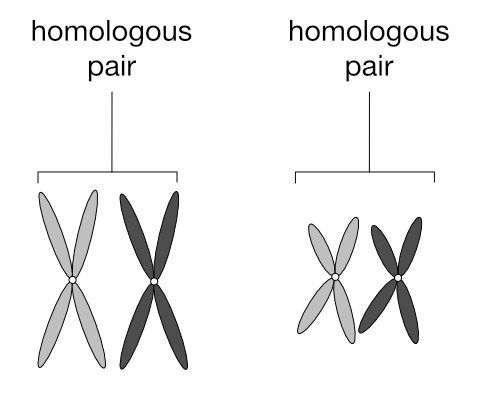
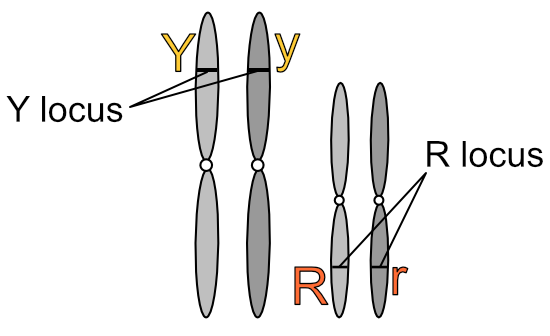
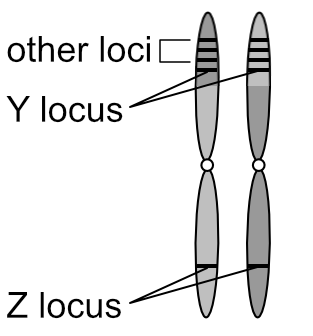
In the 2n cell above, ignore the colours and focus on the two pairs of chromosomes that are similar in shape: the long pair and the short pair. Each of these pairs are called homologous pairs (from the Greek homos, “same” and logos, “relation”). A 2n cell has n pairs of homologous chromosomes. In the diagram below one of each homologous pair is dark grey and the other light grey to make it easier to follow the movements of these pairs during meiosis:
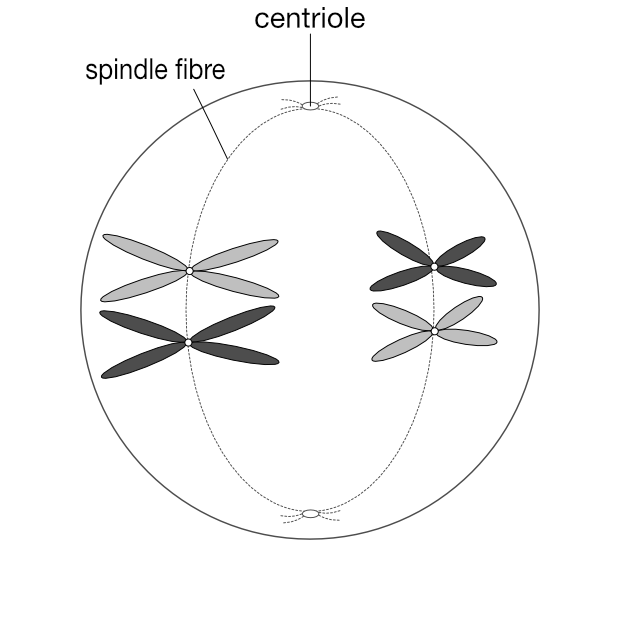
Metaphase 1
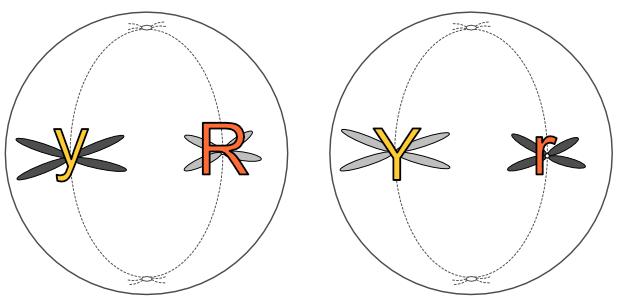
The homologous pairs line up along the equatorial plane of the cell. The nuclear membrane disappears at the end of this stage as it does during mitosis.
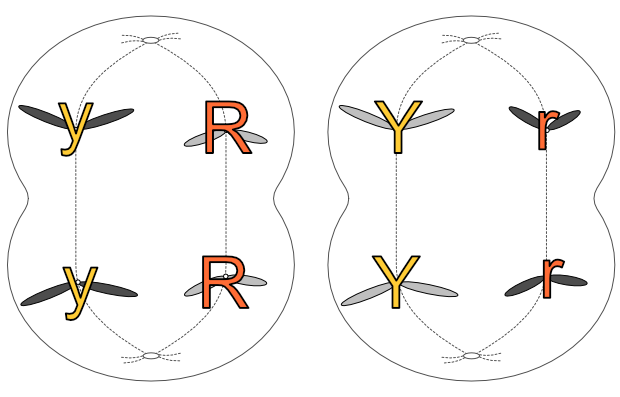
Anaphase 1
There is a crucial distinction here between mitosis and meiosis. The chromatids separate at this stage in mitosis, but in meiosis the chromatids stay together and it is the homologous pairs that separate. it is one of each pair that goes to either pole of the cell in anaphase 1.
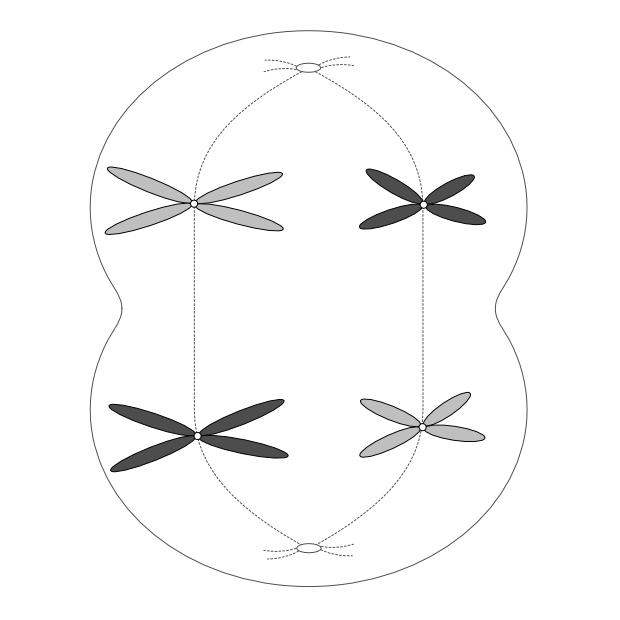
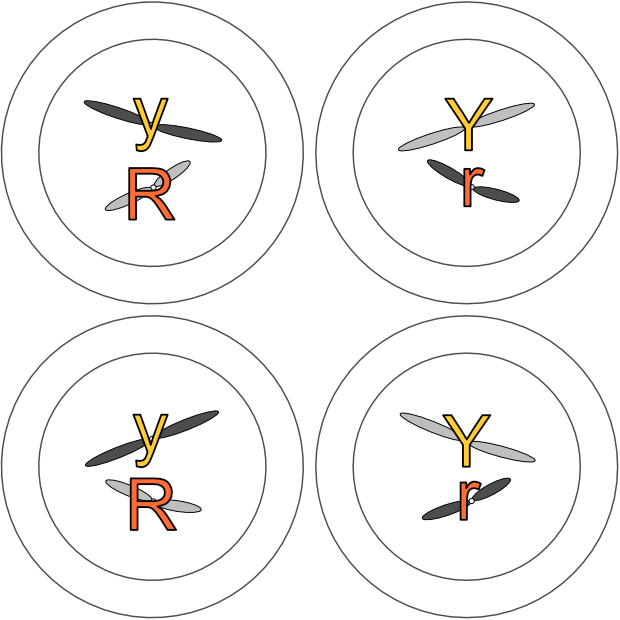
Telophase 1
As in mitosis, a nuclear membrane forms around each group of chromosomes, the cell membrane indents, and two new cells result.
But as you can see from the colour-coding, these cells are not identical as they would be after mitosis. This is another key difference with meiosis at this stage. Homologous chromosomes look the same and carry the same genes, but they do not necessarily carry the same versions of those genes.
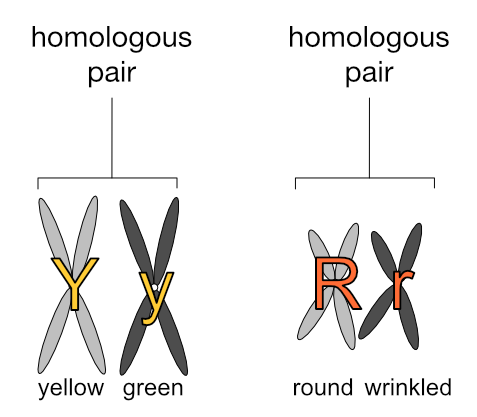
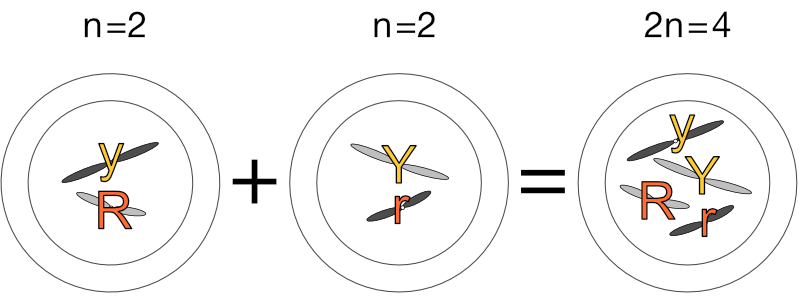
Let’s assign some values to these homologous chromosomes to make this clearer. Going back to Mendel’s peas, let’s have the long chromosomes carry the genes for seed colour (yellow or green) and the short chromosomes carry the genes for seed shape (round or wrinkled). Let’s further have the light grey chromosomes carry the dominant gene (’Y’ or ‘R’) and the dark grey chromosomes carry the recessive gene (’y’ or ‘r’). (Please note this is very simplified and purely for illustration — pea plants have more chromosomes and genes than this.)
We have:
Let’s continue!
Phase 2
The second division. At the end of this stage the chromosomes have separated but have not replicated (they did that in Phase 1).
Interphase 2
This is a very brief stage in Phase 2.
Prophase 2
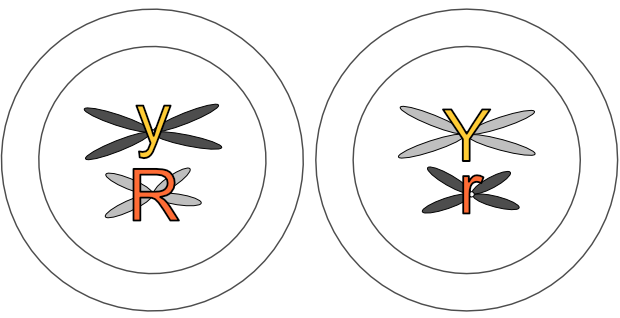
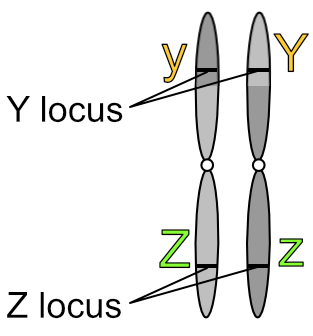
The chromosomes appear as the original pair in telophase 1. Here we’ll continue to use our illustrative pea chromosomes — can you see Mendel’s law of segregation in effect? The ‘Y’ and ‘y’ genes have separated into different cells, as have the ‘R’ and ‘R’ genes.
Metaphase 2
The chromosomes again line up across the middle of each cell:
Anaphase 2
The chromosomes separate and one of each pair goes to opposite ends of the cell.
Telophase 2
The nuclear membrane reforms and the cell divides. The end result is four n cells from one 2n cell.
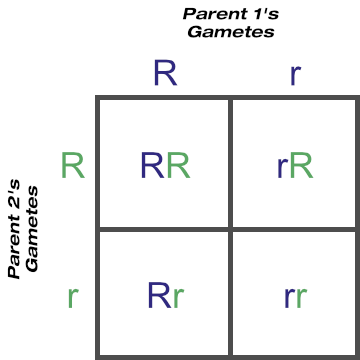
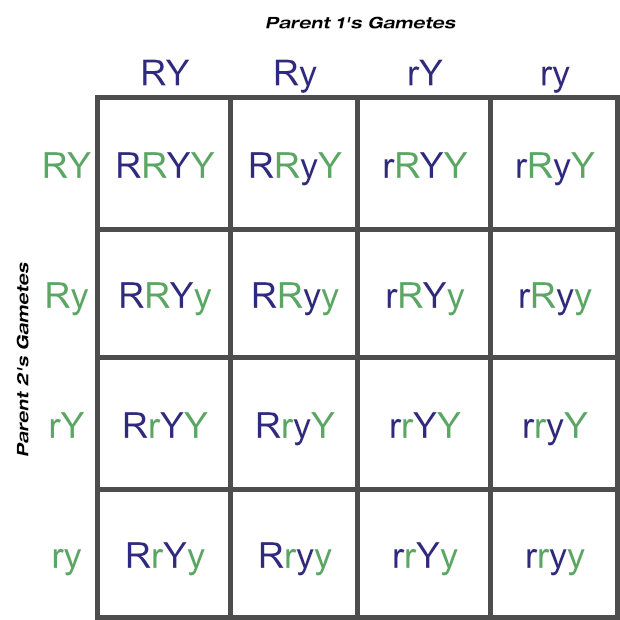
Here we ended up with two Ry cells and two rY cells.
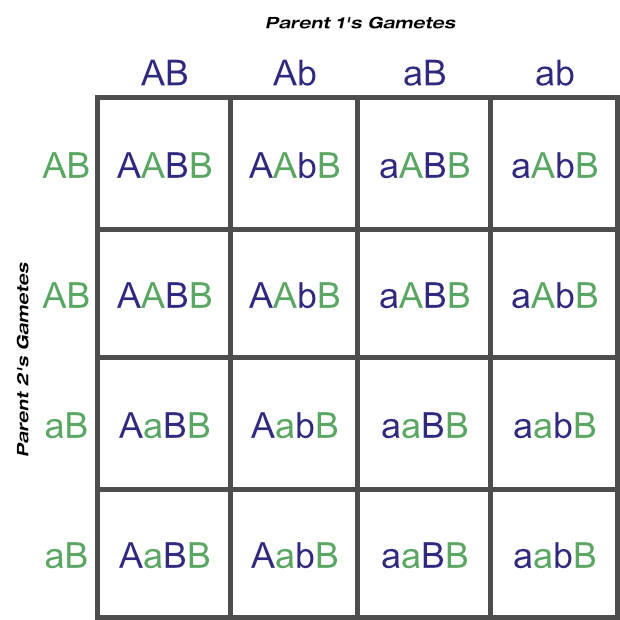
If you go back to the metaphase 1 diagram, you’d see how both light coloured chromosomes could just have easily been aligned at top. Following through the steps, with that scenario we’d have ended up with two RY cells and two ry cells instead. Can you see how Mendel’s law of independent assortment applies here?
And that is meiosis! The example here with four original chromosomes was a very simple one. Imagine the genetic variation in the gametes of dogs (starting with 39 homologous pairs) or cattle (30 homologous pairs)!
We’ve seen how Mendel’s first two laws apply at the cellular level: next we’ll go deeper still into the chromosome and cover Mendel’s third law, the law of dominance. But with a twist, for not all inheritance is Mendelian!